BRFSS Data in R
The CDC's Behavioral Risk Factor Surveillance System (BRFSS) is a "system of health-related telephone surveys that collect state data about U.S. residents regarding their health-related risk behaviors, chronic health conditions, and use of preventive services." The BRFSS was initially established in 1984, and as of 2019 "completes more than 400,000 adult interviews each year, making it the largest continuously conducted health survey system in the world" (CDC 2019).
This tutorial demonstrates how to process and analyze anonymized survey responses in R.
Importing and Processing BRFSS Response Data
Download
Processed BRFSS data is made available in a variety of aggregations and settings. The raw, anonymized survey responses are made available as a fixed width column ASCII file from the CDC website..
Download and unzip the ASCII data file. The 2020 file used in this example was about 44 Mb zipped and 820 Mb unzipped, containing 401,958 rows and 279 columns.
Variable Layout
The information on column positions is available as an HTML table, that you can copy into a CSV file. A version as a CSV is available here.
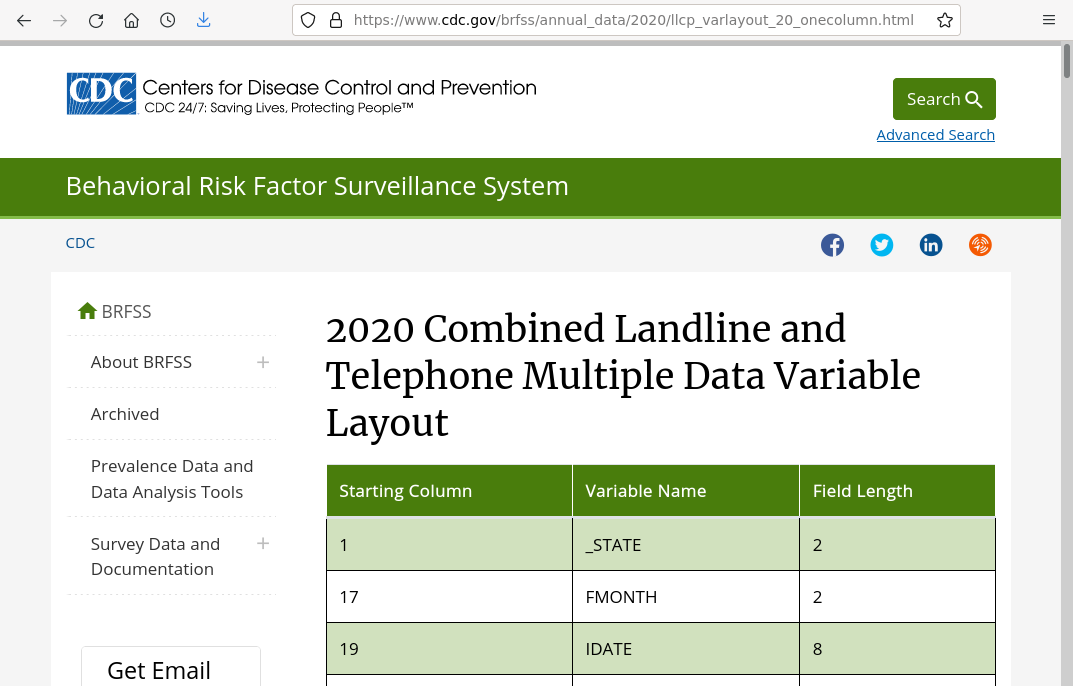
Note that the variable layout includes group fields that encompass multiple subsequent columns and have no independent width of their own (example: IDATE includes IMONTH, IDAY, and IYEAR). There are also blank columns between some fields. This code creates a data frame of columns that removes the group fields and calculates actual field widths in the file (File_Width) for each field, which will be needed to parse the file.
columns = read.csv("2020-column-layout.csv")
columns$File_Width = sapply(1:nrow(columns), function(y) ifelse(y < nrow(columns),
columns$Starting_Column[y + 1] - columns$Starting_Column[y], 1))
columns = columns[columns$File_Width > 0,]
Import File
Given the survey data .txt file and the field widths, you can use the read.fwf() function to import the data into a data frame. That function is a bit slow, and the call took around seven minutes for me. There are apparently faster functions like readr, but since this workflow involves copying out a subset into a new file, the extra effort may not be worth it unless you will be reading this file frequently.
Note that the trailing space in the file name is intentional since the file was apparently zipped with that trailing space.
responses = read.fwf("LLCP2020.ASC ", widths = columns$File_Width, col.names = columns$Variable_Name)
Select Columns
You will probably want to select and modify a specific subset of columns for your uses.
The CDC data page includes a codebook which describes the questions associated with each variable and the range / meaning of the field values.
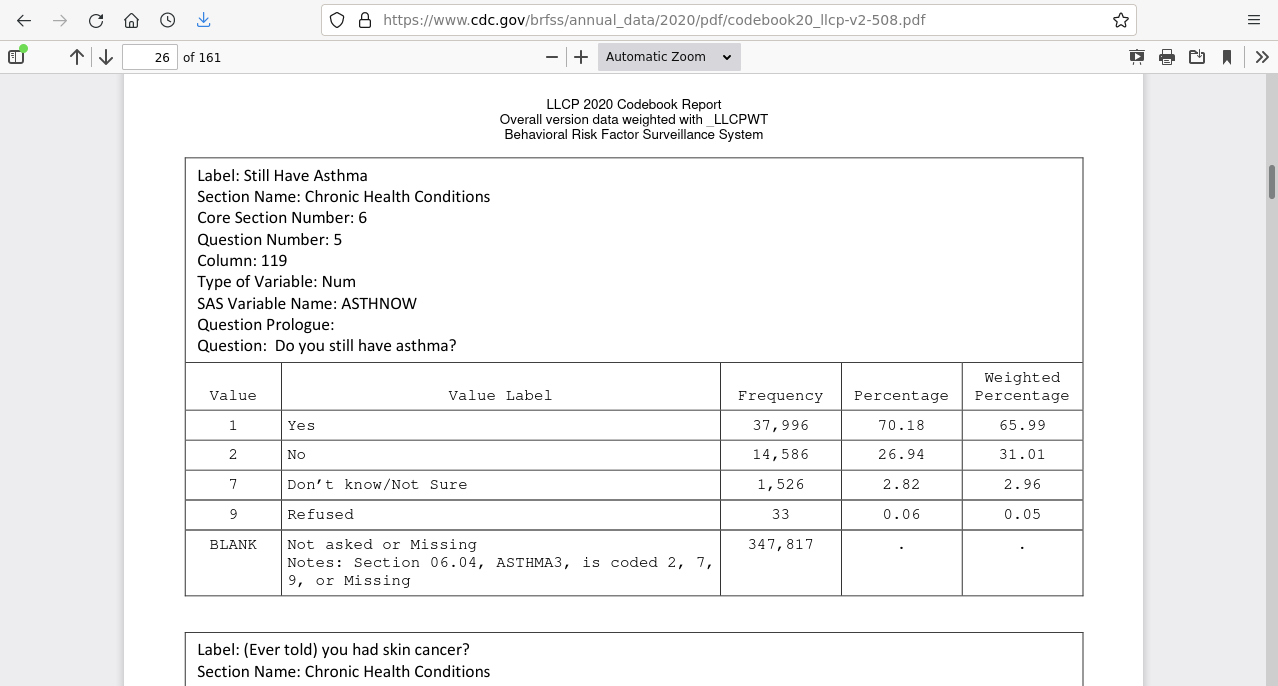
To make the response data more manageable, this code subsets a collection of notable columns with fairly consistent availability across all states.
name_list = c("X_STATE", "X_URBSTAT", "X_AGEG5YR", "DISPCODE", "NUMADULT", "SEXVAR", "GENHLTH",
"PHYSHLTH", "MENTHLTH", "HLTHPLN1", "EXERANY2", "SLEPTIM1", "CVDINFR4",
"CVDCRHD4", "CVDSTRK3", "ASTHMA3", "CHCSCNCR", "CHCOCNCR", "CHCCOPD2",
"HAVARTH4", "ADDEPEV3", "CHCKDNY2", "DIABETE4", "DIABAGE3", "LASTDEN4",
"RMVTETH4", "MARITAL", "EDUCA", "RENTHOM1", "VETERAN3", "EMPLOY1", "CHILDREN",
"INCOME2", "PREGNANT", "WEIGHT2", "HTIN4", "DEAF", "BLIND", "SMOKDAY2",
"STOPSMK2", "AVEDRNK3", "DRNK3GE5", "SEATBELT", "HADMAM", "PSATEST1",
"COLNSCPY", "HIVTST7", "HIVRISK5", "X_LLCPWT")
responses = responses[, name_list]
FIPS Codes
The only specific geospatial field in the data are FIPS (Federal Information Processing Standards) codes for states.
As a convenience, code adds a ST state abbreviation column based on the FIPS codes.
fips = read.csv("fips.csv")
colnames(responses)[colnames(responses) == "X_STATE"] = "FIPS"
responses = merge(fips[,c("FIPS", "ST")], responses, by = "FIPS", all.x=F, all.y=T)
Age Adjustment Weights
Risk for chronic health conditions tends to increase with age. When working with geospatial health data, incidence rates are commonly age-adjusted so that different areas can be compared more meaningfully regardless of whether they have unusually large concentrations of young or old residents (CDC 2021).
The CDC commonly age adjusts based on the proportion of five-year age groups (Klein and Schoenborn 2001). This code adapts 2019 population estimate proportions (USCB 2020) to get an age group weighting vector.
age.weights = c("18-24" = 0.118414, "25-29" = 0.09212, "30-34" = 0.087897,
"35-39" = 0.085178, "40-44" = 0.078063, "45-49" = 0.079928,
"50-54" = 0.08024, "55-59" = 0.085726, "60-64" = 0.080608,
"65-69" = 0.068397, "70-74" = 0.05497, "75-79" = 0.037824,
"80+" = 0.050635, "No Response" = 0)
barplot(age.weights)
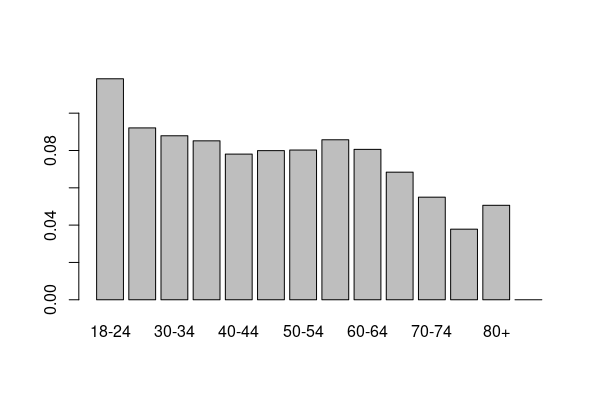
As a convenience, this code adds an X_AAWEIGHT age-adjusted weighting column that is the product of the X_LLCPWT survey weighting column and the age group for each response. This variable can then be used for weighting when calculating age-adjusted means and proportions.
responses$X_AAWEIGHT = NA
for (state in na.omit(unique(responses$ST))) {
state_rows = responses$ST %in% state
groups = table(responses[state_rows, "X_AGEG5YR"])
groups = groups / sum(groups)
groups = age.weights[1:length(groups)] / groups
responses[state_rows, "X_AAWEIGHT"] =
responses[state_rows, "X_LLCPWT"] *
groups[responses[state_rows, "X_AGEG5YR"]]
}
Save CSV
Once you have processed the fixed-width file into a data frame that fits your needs, you may find it valuable to save that data frame to a CSV (comma-separated variable) file so you can save the processing time if you need the data in the future.
write.csv(responses, "2020-brfss-survey-responses.csv", row.names=F);
Import
A CSV file of 2020 data processed using the steps above is available for download here.
Tabular data can be read into a data frame using the read.csv() file. Data frames are analogous to spreadsheets with columns of data for variables arranged in rows.
responses = read.csv("2020-brfss-survey-responses.csv")
The dim() function shows the dimensions of the data frame in number of rows (survey responses in this example) and number of columns (variables on each response in this example).
dim(responses)
[1] 401958 49
The names() function shows the available variables. Consult the metadata (or the BRFSS codebook linked above) for documentation of what the variables represent and what the ranges of values are.
names(responses)
[1] "FIPS" "X_URBSTAT" "X_AGEG5YR" "DISPCODE" "NUMADULT" "SEXVAR" [7] "GENHLTH" "PHYSHLTH" "MENTHLTH" "HLTHPLN1" "EXERANY2" "SLEPTIM1" [13] "CVDINFR4" "CVDCRHD4" "CVDSTRK3" "ASTHMA3" "CHCSCNCR" "CHCOCNCR" [19] "CHCCOPD2" "HAVARTH4" "ADDEPEV3" "CHCKDNY2" "DIABETE4" "DIABAGE3" [25] "LASTDEN4" "RMVTETH4" "MARITAL" "EDUCA" "RENTHOM1" "VETERAN3" [31] "EMPLOY1" "CHILDREN" "INCOME2" "PREGNANT" "WEIGHT2" "HTIN4" [37] "DEAF" "BLIND" "SMOKDAY2" "STOPSMK2" "AVEDRNK3" "DRNK3GE5" [43] "SEATBELT" "HADMAM" "PSATEST1" "COLNSCPY" "HIVTST7" "HIVRISK5" [49] "ST"
Subsetting
Imported data will commonly contain missing values (NA) or special codes that need to be subsetted out in order to use the data meaningfully.
Data frames can be treated like arrays and subsets are taken by following the data frame name with square brackets.
- The first index value in the brackets select the rows and the second index value selects the columns. The two values are separated by a comma.
- Row selection conditions are usually logical values created with comparisons (<, <=, ==, >=, >). Multiple logical conditions can be combined with the AND (&) or OR (|) operators.
- Columns selection conditions are usually column names. Multiple columns can be selected by including the names in the c() concatenation function.
- If either the first or second index value is blank, all rows or columns, respectively, are selected.
For example, with the HTIN4 (height in inches) variable used in this example, the only cleaning needed will be removal of blank (NA) values:
height = responses[!is.na(responses$HTIN4),] quantile(height$HTIN4, na.rm=T)
0% 25% 50% 75% 100% 36 64 67 70 95
Special Codes
This particular BRFSS data set often mixes special codes in with quantitative values. Subsetting can remove those codes so you have only the quantitative values.
For example, the SLEPTIM1 (hours of sleep per 24-hour period) uses the values 77 for "Don't know/Not Sure" and 99 for "Refused," as well as NA (blank) if the data is not available.
sleep = responses[responses$SLEPTIM1 %in% 1:24, ] quantile(sleep$SLEPTIM1)
0% 25% 50% 75% 100% 1 6 7 8 24
WEIGHT2 (weight) is more complex, with codes for don't know / refused and specific ranges for respondents that answer in kilograms rather than pounds. The codes can be removed along with the NA values by subsetting values that are within the possible pound range of 50 to 776(!).
weight = responses[responses$WEIGHT2 %in% 50:776,] quantile(weight$WEIGHT2)
0% 25% 50% 75% 100% 50 150 175 205 758
For some variables, the code 88 is used for zero to more clearly separate nonzero values of interest. Those values can be selectively changed to zero with subsetting.
health = responses[responses$PHYSHLTH %in% c(1:30, 88),] health[health$PHYSHLTH == 88, "PHYSHLTH"] = 0 quantile(health$PHYSHLTH, na.rm=T)
0% 25% 50% 75% 100% 0 0 0 2 30
Distributions
A frequency distribution is "an arrangement of statistical data that exhibits the frequency of the occurrence of the values of a variable" (Merriam-Webster 2022).
The type of distribution will dictate what kinds of statistical techniques you should use on your data.
Frequency Tables
For categorical variables, the table() function can be used to tally frequency for each value in the distribution.
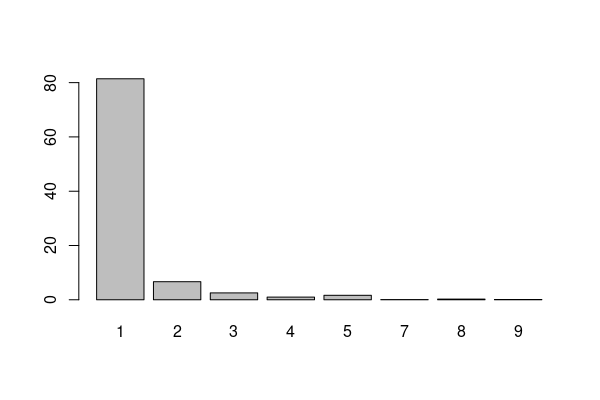
The mode for this distribution is clearly 1 (always wear a seat belt).
seatbelts = table(responses$SEATBELT, useNA="no") print(seatbelts)
1 2 3 4 5 7 8 9
327249 26763 10195 3942 6632 263 976 427
Dividing the table by the number of responses and multiplying by 100 gives percentages that can be visualized as a bar graph with the barplot() function.
barplot(seatbelts * 100 / nrow(responses))
Histograms
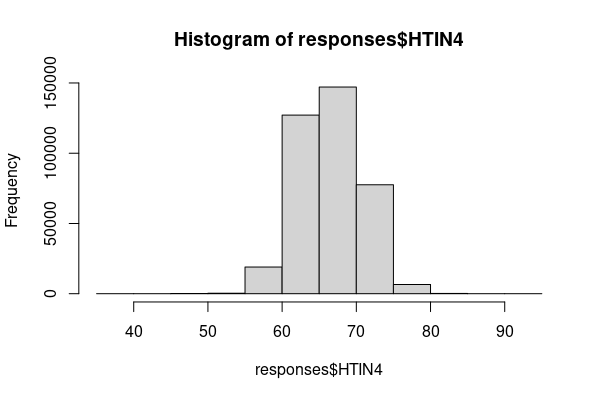
For quantitative variables, a fundamental way of visualizing the frequency distribution is the histogram, which is a bar graph that shows the number of values within different ranges of values.
In R, the hist() function can be used to create a quick histogram. In this data, the HTIN4 variable is height in inches.
height = responses[!is.na(responses$HTIN4),] hist(height$HTIN4)
Density Plots
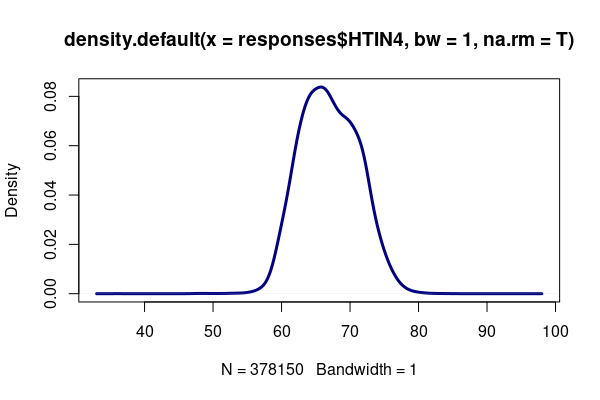
An alternative visualization to the histogram is a density plot that shows clearer detail by smoothing the histogram in manner similar to a continuous density distribution.
height = responses[!is.na(responses$HTIN4),] plot(density(height$HTIN4, bw=1, na.rm=T), lwd=3, col="navy")
Normal Distributions
A normal distribution is an extremely common distribution in both the social and environmental sciences where data tends to cluster symmetrically around a central mean value with proportions of values decreasing as you move further above or below the mean. This distribution deserves special attention because many statistical techniques assumes variables that are normally distributed, and you should use caution in selecting analytical methods to make sure your method is appropriate for your distribution.
The frequency distribution curve for the normal distribution is often called the bell curve because of its bell-like shape. The density formula for that curve is a bit ugly:
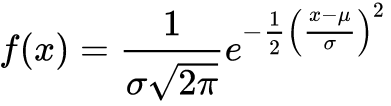
R contains functions for analyzing and simulating a variety of distributions. The dnorm() function returns the density curve value for a specific value within the curve, and is used here to graph the normal curve:
plot(dnorm(seq(-4, 4, 0.1)), type="l", lwd=3, col="navy")
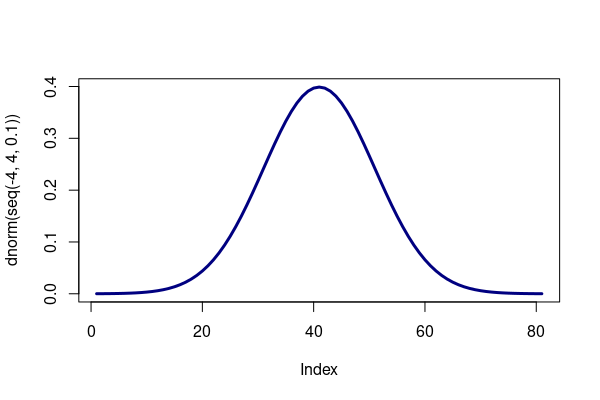
The height variable roughly follows a normal distribution, in which values tend to cluster around a central (mean) value, with fewer values as you get further above or below the mean.
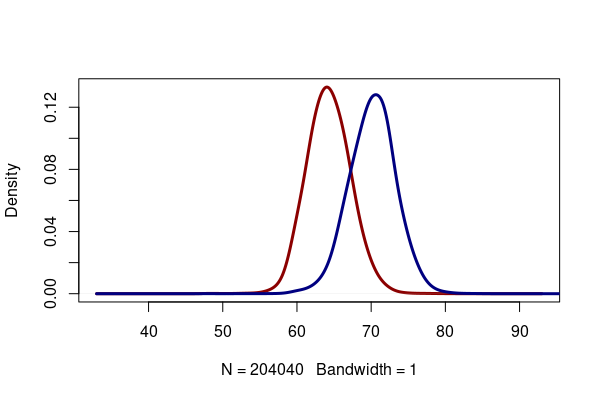
In the height density curve above, you may notice a slight notch on the right side of the curve. This is because men and women have separate normal distributions of height, and adding them together creates an asymmetrical curve.
height = responses[!is.na(responses$HTIN4),] men = height[height$SEXVAR == "1", "HTIN4"] women = height[height$SEXVAR == "2", "HTIN4"] plot(density(women, bw=1, na.rm=T), lwd=3, col="darkred", main="") lines(density(men, bw=1, na.rm=T), lwd=3, col="navy")
Skew
A normal distribution is symmetrical in that there are an equal number of values above and below a central value, and the values change in a similar manner as you move above or below that central value.
Skew occurs when the distribution contains values extend a greater distance from the mean either on the high (right skew) or low side (left skew).
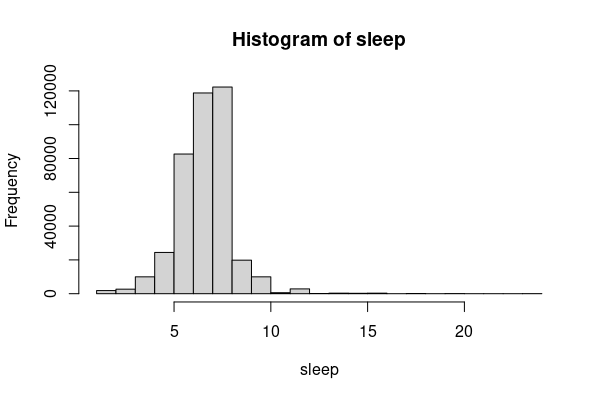
For example in this dataset, the number of hours that most people sleep per night is around seven hours per night, but there are a limited number of people who claim to sleep considerably more that, which skews the right tail of the curve far from the mean.
sleep = responses[responses$SLEPTIM1 %in% 1:24, ] hist(sleep$SLEPTIM1)
Modality
The mode of a distribution is the most common value in the distribution. With quantitative variables, the mode is the peak of the density curve.
Distributions like the normal distribution are unimodal because they have one peak value around which a smaller number of higher and lower values cluster.
Multimodal distributions have more than one prominent peak value.
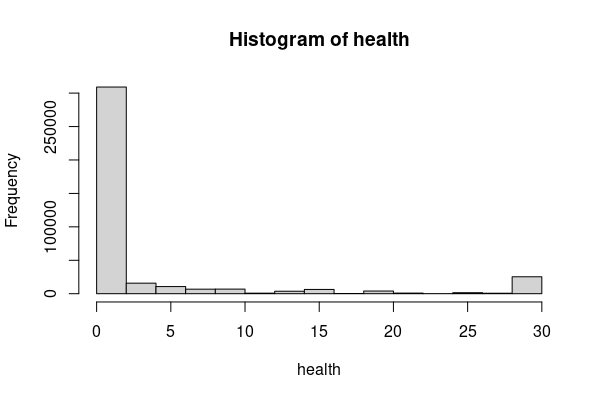
For example in this data, the number of days over the past 30 days that respondents indicated that they felt their physical health was not good clusters into two modes representing generally well people (zero on one day) and people with chronic illnesses (all 30 days).
health = responses[responses$PHYSHLTH %in% c(1:30, 88),] health[health$PHYSHLTH == 88, "PHYSHLTH"] = 0 hist(health$PHYSHLTH)
Measures of Central Tendency
When working with a variable, we often wish to summarize the variable with a measure of central tendency, which is a single value that represents the center of the distribution.
There are three primary types of central tendency values: mean, median, and mode.
Mean
The arithmetic mean of a variable is the sum of all values divided by the number of values. In common parlance, arithmetic means are often simply called means or averages.
In R, the mean() function calculates the arithmetic mean of the vector or field passed to it. The na.rm=T parameter is often helpful since mean() will return NA if any values within the variable are NA, and na.rm=T instructs mean() to ignore NA values.
height = responses[!is.na(responses$HTIN4),] plot(density(height$HTIN4, bw=1, na.rm=T), lwd=3, col="navy", main="") x = mean(height$HTIN4, na.rm=T) abline(v=x, col="darkred", lwd=3) print(x)
[1] 66.99402
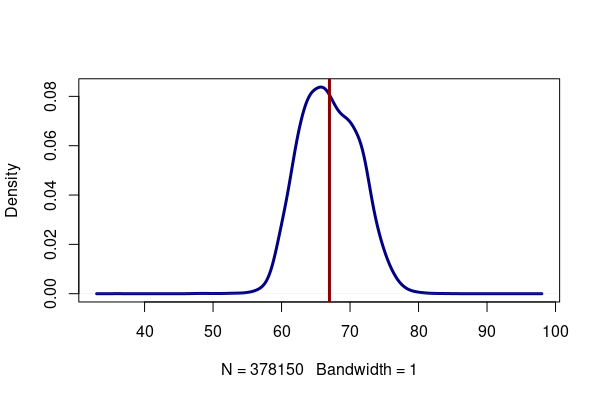
Median
A median is a central tendency value where half of the values in the distribution are below the median and half are above the median.
Medians are often used rather than means with non-normal or heavily skewed distributions where outliers can make the mean unrepresentative of the phenomenon being summarized.
For example, if a billionaire developer walks into a room full of construction workers, the mean income will jump dramatically, while the median will stay largely the same and be more representative of the income of the people in the room.
In a symmetrical distribution like the normal distribution, the mean and median should be nearly identical.
In skewed distributions, the mean will be biased in the direction of the outliers. Whether that is undesirable depends on what you wish to convey in your analysis. For example, with the skewed distribution of sleep hours per night, the median is an even seven, which reflects the bulk of the responses. But the mean is slightly higher, which more-clearly communicates that there are some folks who get more than seven hours of sleep per night.
sleep = responses[responses$SLEPTIM1 %in% 1:24, ]
sleep.mean = mean(sleep$SLEPTIM1, na.rm=T)
sleep.median = median(sleep$SLEPTIM1, na.rm=T)
plot(density(sleep$SLEPTIM1, na.rm=T, bw=0.5), col="gray", lwd=3, main="")
abline(v=sleep.mean, col="navy", lwd=3)
abline(v=sleep.median, col="darkred", lwd=3)
legend("topright", lwd=3, lty=1, col=c("navy", "darkred"), legend=c("Mean", "Median"))
print(sleep.mean)
print(sleep.median)
[1] 7.097099 [1] 7
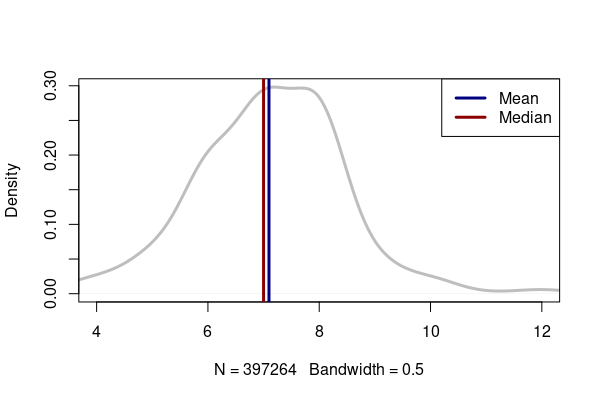
Mode
The mode of a distribution is the most common value in the distribution.
This concept is most clearly applicable categorical variables that have a limited number of possible values, such Likert scales that convert qualitative perspectives into ordinal quantitative values.
For example, the SEATBELT question is a Likert scale based on respondent ratings of their seatbelt usage on a scale of one (always) to five (never). Additional codes are included for people who were unsure, who never drive, or who refused to answer the question.
The table() function tallies frequency for each value in the distribution.
The mode for this distribution is clearly 1 (always wear a seat belt).
seatbelts = table(responses$SEATBELT) print(seatbelts)
1 2 3 4 5 7 8 9
327249 26763 10195 3942 6632 263 976 427
Dividing the table by the number of responses and multiplying by 100 gives percentages that can be visualized as a bar graph with the barplot() function.
barplot(seatbelts * 100 / nrow(responses))

While this visualization shows a clear dominance of people who respond that they always wear seat belts, keep in mind that with responses that involve politically-sensitive issues or imply a moral judgment, people often exhibit social desirability bias.
Bias is "systematic error introduced into sampling or testing by selecting or encouraging one outcome or answer over others" (Merriam-Webster 2022).
Social desirability bias is a problem with surveys in which participants (intentionally or unintentionally) respond with the answer they think they are supposed to give rather than an answer that accurately reflects what they actually do in their lives (Krumpal 2013).
Measures of Variation
In order to fully describe a distribution you need not only a central value, but a measure of how the values in a distribution vary across the range of values.
Range
The simplest measure of variation with a quantitative value is the range() of values.
Note that ranges tell you nothing about the frequency of values, so outliers can cause range to give a distorted picture of what the data actually shows.
height = responses[!is.na(responses$HTIN4),] range(height$HTIN4, na.rm=T)
[1] 36 95
For categorical variables, the unique() function shows all possible values.
unique(responses$MARITAL)
[1] 3 1 2 4 5 9 6 NA
Percentiles
A percentile is "a value on a scale of 100 that indicates the percent of a distribution that is equal to or below it" (Merriam-Webster 2022).
- The zero percentile is the lowest value in the distribution.
- The 50th percentile is the median (the middle value in the distribution).
- The 100th percentile is the highest value in the distribution.
Percentiles can be calculated by passing a probability in the prob parameter of the quantile() function.
For example, this code uses the quantile() function to calculates the 25th percentile of height, which means that 25% of male respondents were at or below 68 inches in height.
height = responses[!is.na(responses$HTIN4),] men = height[height$SEXVAR %in% 1, ] quantile(men$HTIN4, prob=0.25)
25% 68
Going the other direction and asking what percent of respondents meet a condition requires a bit of calculation based on the number of rows in a subset that meet that condition, divided by the total number of rows, and multiplied by 100 to get a percentage.
over.six.feet = 100 * sum(men$HTIN4 > 72) / nrow(men) paste0(round(over.six.feet,2) , "% of male respondents are over 6 feet tall.")
[1] "20.07% of male respondents are over 6 feet tall."
Quantiles
A quantile is a set of values that divide a frequency distribution into classes (bins) that each contain the same percentage of the total population.
The values dividing bins are percentiles. Quantiles are especially useful for describing non-normal distributions.
Quantiles with commonly used numbers of classes have their own names. A quartile is three percentile values that divide a distribution into four groups. A quintile is four percentile values that divide a distribution into five groups.
The quantile() function can display quantile values for a variable. By default, the quantiles are displayed in 25% percentile intervals with the 0% percentile being the lowest value in the range, the 50% percentile being the median, and the 100% percentile being the highest value in the range.
height = responses[!is.na(responses$HTIN4),] quantile(height$HTIN4)
0% 25% 50% 75% 100% 36 64 67 70 95
A barplot() can be used to visualize a quantile(), although such graphics can be confused with histograms.
barplot(quantile(height$HTIN4))
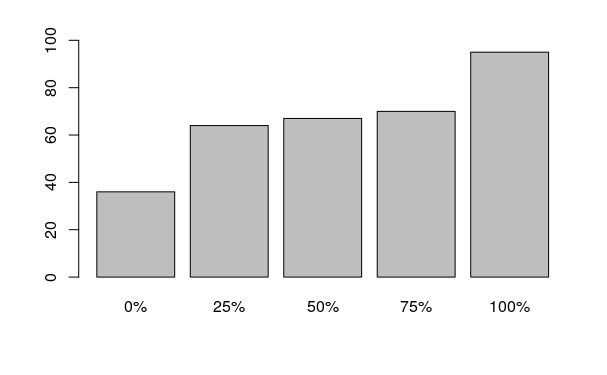
Quantiles comparing different groups are commonly visualized as side-by-side box-and-whisker plots using the boxplot() functions. By default, box-and-whisker plots in R represent the following values (Ford 2015).
- The line through the middle is the median.
- The gray box outlines the 25th to 75th percentile.
- The whiskers outline 1.5 times the interquartile range (IQR), which is the difference between the 25th and 75th percentile.
- The dots outside the whiskers are outliers.
height = responses[!is.na(responses$HTIN4) & !is.na(responses$SEXVAR),] boxplot(HTIN4 ~ SEXVAR, data = height)
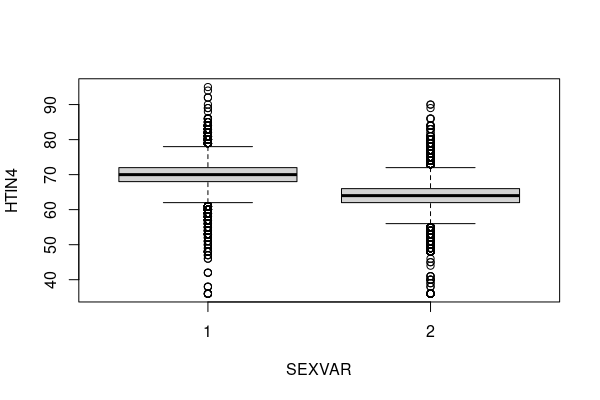
Standard Deviation
With normally-distributed data, the width of the bell curve is specified by the standard deviation. Standard deviation is represented by the lower-case Greek letter sigma (σ), which looks like a lower-case English "b" that's had too much to drink. Standard deviation for discrete values is calculated as the square root of the sums of the squares of deviations of the values (xi) from the mean (μ).
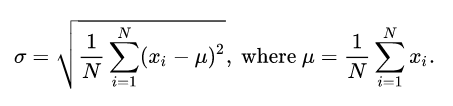
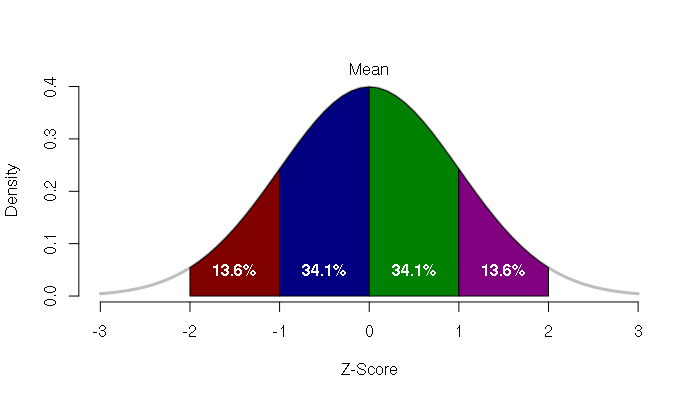
The number of standard deviations that a value departs from the mean is called the z-score. The z-score of the mean is zero.
Converting values to z-scores is called standardizing, which allows comparison between values from different distributions.
The area under the bell curve between a pair of z-scores gives the percentage of things associated with that range range of values. For example, the area between one standard deviation below the mean and one standard deviation above the mean represents around 68.2 percent of the values.
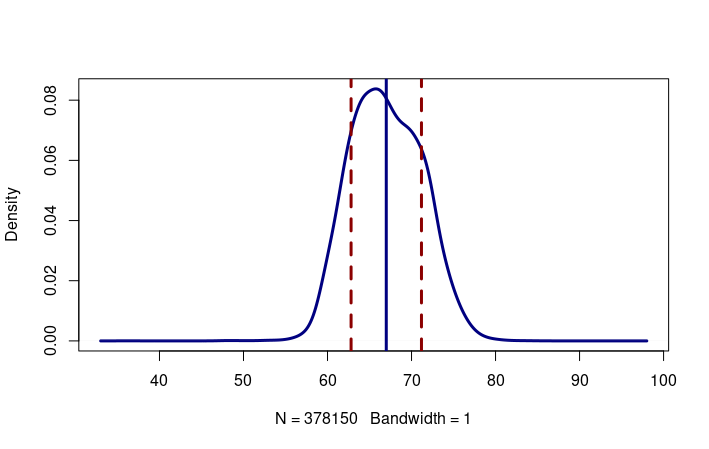
The sd() function can be used to calculate the standard deviation for a set of values. For example, this plots the density curve for height, along with vertical lines for the mean, one standard deviation below the mean (z = -1) and one standard deviation above the mean (z = 1).
height = responses[!is.na(responses$HTIN4),]
height.mean = mean(height$HTIN4, na.rm=T)
height.sd = sd(height$HTIN4, na.rm=T)
plot(density(height$HTIN4, bw=1, na.rm=T), lwd=3, col="navy", main="")
abline(v=height.mean, col="navy", lwd=3)
abline(v=height.mean - height.sd, col="darkred", lty=2, lwd=3)
abline(v=height.mean + height.sd, col="darkred", lty=2, lwd=3)
paste("The mean US height is", height.mean,
"inches with a standard deviation of",
height.sd, "inches.")
The mean US height is 66.99402 inches with a standard deviation of 4.188529 inches.
Coefficient of Variation
One challenge with the standard deviation is that it is an absolute value that needs to be assessed in the context of the mean, which makes it difficult to compare the standard deviations for two different distributions.
One solution is to use the coefficient of variation. The coefficient of variation is the standard deviation divided by the mean, which gives a value that represents a percentage variation.
For example, with our height data, the coefficient of variation for women is slightly higher than that for men, which indicates that women's heights vary slightly more than men's heights.
height = responses[!is.na(responses$HTIN4),]
height.men = height[height$SEXVAR == 1,]
height.women = height[height$SEXVAR == 2,]
covar.men = sd(height.men$HTIN4) / mean(height.men$HTIN4)
covar.women = sd(height.women$HTIN4) / mean(height.women$HTIN4)
paste0("Coefficient of variation: men = ",
round(100 * covar.men, 2),
"%, women = ",
round(100 * covar.women, 2), "%")
[1] "Covariance: men = 4.42%, women = 4.51%"
Skew and Kurtosis
Skew is the extent to which values are evenly distributed around the mean. The skewness() function from the e1071 library can be used to calculate a value that will be negative if the left tail of the distribution is longer, and positive if the right tail of the distribution is longer. The skewness of a normal distribution is zero.
Kurtosis is a measurement of how flat or peaked the distribution is. The kurtosis() function from the e1071 library can be used to calculate a value that is positive for a sharply peaked distribution and negative for a distribution that is flatter than a normal distribution. The kurtosis of a normal distribution is zero.
For the example skewed distribution of hours of sleep per night, the positive skewness() corresponds to the long right tail and the high kurtosis() corresponds to the sharp peak around seven hours / night.
library(e1071) sleep = responses[responses$SLEPTIM1 %in% 1:24,] print(skewness(sleep$SLEPTIM1)) print(kurtosis(sleep$SLEPTIM1))
[1] 0.7551364 [1] 8.574684
In contrast, the largely normal distribution for height has very small values of both skewness and kurtosis.
height = responses[!is.na(responses$HTIN4),] print(skewness(height$HTIN4)) print(kurtosis(height$HTIN4))
[1] 0.08109344 [1] -0.08372729
Weighting
The downloadable BRFSS data is raw, anonymized survey data that needs to be weighted because it is biased by uneven geographic coverage of survey administration (noncoverage) and lack of responsiveness from some segments of the population (nonresponse).
The X_LLCPWT field (landline, cellphone weighting) is a weighting factor added by the CDC that can be assigned to each response to compensate for these biases. See Weighting the BRFSS Data (CDC 2020) for more details on the calculation of these weighting factors.
X_LLCPWT can be used for weighting when calculating crude means and proportions at the state or national level.
The X_AAWEIGHT field is an age-adjusted weighting field (created above) that can be used for calculating age-adjusted means and proportions at the state or national level.
Weighted Means
The BRFSS data contains a handful of quantitative variables, although some assess comparatively rare behaviors or conditions that result in highly skewed distributions. In addition, some fields add special codes or ranges that need to be handled.
In these examples:
- Raw rates are simple means of the sample data with no weighting.
- Crude rates are means weighted by the CDC weighting factor that compensates for noncoverage and nonresponse.
- Age-adjusted rates are crude rates that are further weighted to compensate for areas with unusually high or low percentages of different age groups.
HTIN4 (height in inches) has no special codes and can be used after removing NA values and NA weights.
height = responses[!is.na(responses$HTIN4) & !is.na(responses$X_AAWEIGHT),] height.raw = mean(height$HTIN4) height.crude = weighted.mean(height$HTIN4, height$X_LLCPWT) height.age.adjusted = weighted.mean(height$HTIN4, height$X_AAWEIGHT) print(height.raw) print(height.crude) print(height.age.adjusted)
[1] 67.02791 [1] 67.04312 [1] 67.20037
WEIGHT2 (weight) is more complex, with codes for don't know / refused and specific ranges for respondents that answer in kilograms rather than pounds. The codes can be removed along with the NA values by subsetting values under the maximum pound range of 776(!).
weight = responses[(responses$WEIGHT2 %in% 50:776) & !is.na(responses$X_AAWEIGHT),] weight.raw = mean(weight$WEIGHT2) weight.crude = weighted.mean(weight$WEIGHT2, weight$X_LLCPWT) weight.age.adjusted = weighted.mean(weight$WEIGHT2, weight$X_AAWEIGHT) print(weight.raw) print(weight.crude) print(weight.age.adjusted)
[1] 181.7209 [1] 181.0673 [1] 180.6846
A comparative density plot shows the age adjusted body weight distribution shifted slightly lower than the distribution of raw samples.
plot(density(weight$WEIGHT2, weights=weight$X_AAWEIGHT /
sum(weight$X_AAWEIGHT), bw=5), lwd=3, col="navy",
xlim=c(0,400), main="")
lines(density(weight$WEIGHT2, bw=5), lwd=3, col="red")
legend("topright", lwd=3, col=c("navy", "red"), legend=c("Age-Adjusted", "Raw"))
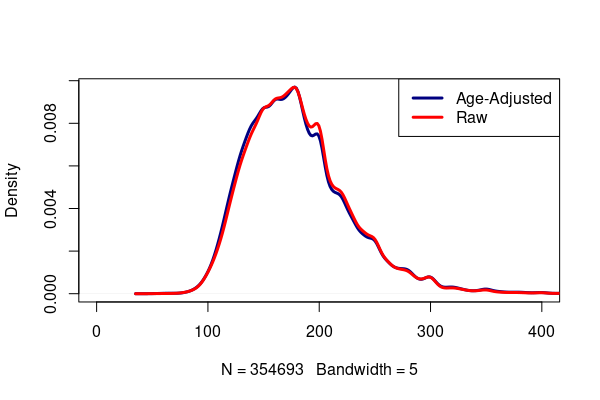
Weighted Mean MOE
Because the BRFSS is a sample, there is a margin of error associated with means. By using the wtd.var() function from the Hmisc library, we can calculate a weighted standard deviation and use that to calculate the 95% confidence interval.
library(Hmisc)
height = responses[!is.na(responses$HTIN4) & !is.na(responses$X_LLCPWT),]
height.mean = weighted.mean(height$HTIN4, height$X_LLCPWT)
height.sd = sqrt(wtd.var(height$HTIN4, height$X_LLCPWT))
height.stderr = height.sd / sqrt(nrow(height))
height.moe = 1.96 * height.stderr
paste("The mean height is",
round(height.mean, 2), "±",
round(height.moe, 2), "inches")
[1] "The mean height is 67.05 ± 0.01 inches"
Weighted Proportions
Proportions of responses to categorical questions can be weighted by summing the weights in the different categories and dividing by the overall sum of the weights. Multiplying by 100 gives values displayed as percents.
This code shows the crude national rate for heart attacks of around 4.2%. One is yes, two is no, seven is don't know, and nine is no response.
library(weights) heart.percent = wpct(responses$CVDINFR4, responses$X_LLCPWT) print(round(100 * heart.percent, 2))
1 2 7 9
4.20 95.23 0.49 0.08
We can subset the responses to get a specific state.
heart = responses[responses$ST %in% "IL",] heart.il = wpct(heart$CVDINFR4, heart$X_LLCPWT) print(round(100 * heart.il, 2))
1 2 7 9
3.90 95.69 0.26 0.15
By using the X_AAWEIGHT age-adjusted weighting parameter, we can get an age-adjusted rate. Illinois has median age slightly above the US median, which may account for an age-adjusted heart attack rate lower than the crude heart attack rate.
heart = responses[responses$ST %in% "IL",] heart.il = wpct(heart$CVDINFR4, heart$X_AAWEIGHT) print(round(100 * heart.il, 2))
1 2 7 9
3.19 96.41 0.24 0.16
Weighted Proportion MOE
Using the general formula for proportions, we can find the margin of error.
heart.attack = heart.il[1]
heart.stderr = sqrt(heart.attack * (1 - heart.attack) / nrow(heart))
heart.moe = 1.96 * heart.stderr
paste(round(heart.attack * 100, 2), "±",
round(100 * heart.moe, 2),
"percent of IL residents have had heart attacks.")
[1] "3.19 ± 0.57 percent of IL residents have had heart attacks."
Weighted Likert Scales
A Likert scale is a technique commonly used on surveys to scale qualitative responses into ordinal values. Likert scale responses can be treated both as quantitative and categorical variables, although you will need to remove the special codes to get means.
For example, the GENHLTH variable is based on the question, "Would you say that in general your health is..." and respondents can then rank on a scale of one (excellent) to five (poor), with seven for don't know and nine for refused to answer.
- A weighted table of proportions can be created with the wpct() function from the weights library.
- We multiply by 100 to convert proportions to percents and round() to one decimal point to improve readability.
- The vector can also be plotted with barplot().
library(weights) health = responses[responses$ST %in% "IL",] percents = wpct(health$GENHLTH, health$X_AAWEIGHT) print(round(percents * 100, 1)) barplot(percents * 100)
1 2 3 4 5 7 9 24.2 35.0 28.7 9.2 2.7 0.1 0.0
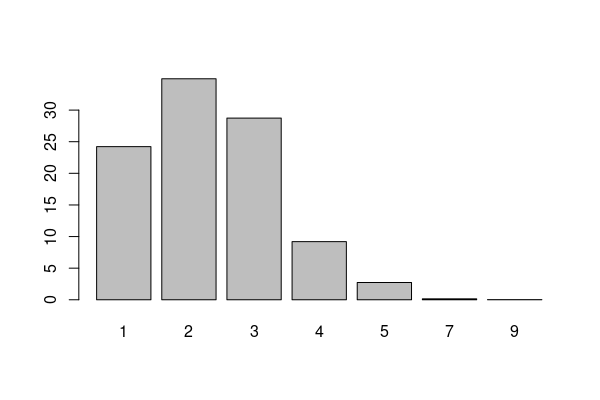
To take a mean of a Likert scale, you should remove or transform the special codes.
health = responses[(responses$GENHLTH %in% 1:5) & !is.na(responses$X_AAWEIGHT),] health = health[health$ST %in% "IL",] health.age.adjusted = weighted.mean(health$GENHLTH, health$X_AAWEIGHT) print(health.age.adjusted)
[1] 2.311411
Weighted Contingency Tables
Contingency tables can be used to detect relationships between categorical variables.
For this example we use SMOKDAY2 (current or former smoker) and CVDINFR4 (heart attack). Note that people who have never smoked have SMOKDAY2 as NA.
smoke = responses[!is.na(responses$X_AAWEIGHT) &
!is.na(responses$CVDINFR4) &
!is.na(responses$SMOKDAY2),]
smoke = aggregate(smoke$X_AAWEIGHT, by=list(Smoker = smoke$SMOKDAY2, Heart.Attack = smoke$CVDINFR4), FUN=sum)
names(smoke)[3] = "Percent"
smoke$Percent = round(100 * smoke$Percent / sum(smoke$Percent), 2)
smoke = reshape(smoke, timevar="Smoker", idvar="Heart.Attack", direction="wide")
names(smoke) = c("Heart.Attack", "Daily", "Some Days", "Never", "Don't Know", "Refused")
print(smoke)
Heart.Attack Daily Some Days Never Don't Know Refused 1 1 1.51 0.51 3.34 0.00 0.01 6 2 26.93 12.10 54.80 0.08 0.16 11 7 0.18 0.06 0.28 0.00 0.00 16 9 0.01 0.00 0.03 NA 0.00
Maps
Mapping Means
Aggregate means " to collect or gather into a mass or whole" (Merriam-Webster 2022).
The aggregate() function can be used to aggregate values by geographic areas.
- The FUN parameter indicates what function is used to aggregate values in each geographic area.
- We sum both the weighted WEIGHT2 and the X_AAWEIGHT age-adjusted weights for each state.
- The state averages can then be calculated by dividing the sum of the weighted values by the sum of the weights.
- Using order() to order the states by weights allows us to list the states with the highest and lowest average weights.
weight = responses[(responses$WEIGHT2 %in% 50:776) & !is.na(responses$X_AAWEIGHT),]
weight$WEIGHT.SUM = weight$WEIGHT2 * weight$X_AAWEIGHT
weight.states = aggregate(weight[,c("WEIGHT.SUM", "X_AAWEIGHT")], by=list(ST = weight$ST), FUN=sum)
weight.states$AVGWEIGHT = weight.states$WEIGHT.SUM / weight.states$X_AAWEIGHT
weight.states = weight.states[order(weight.states$AVGWEIGHT),]
head(weight.states)
ST WEIGHT.SUM X_AAWEIGHT AVGWEIGHT 12 HI 193820169 1147803.9 168.8617 8 DC 117999411 690173.2 170.9707 20 MA 917219772 5297312.6 173.1481 35 NY 2610813502 15026814.8 173.7436 6 CO 806333502 4628338.2 174.2166 32 NJ 1116181092 6387053.7 174.7568
In order to map the aggregated data, we need to join the table data to geospatial polygons that indicate where to draw the features.
A join is an operation where two data frames are combined based on common key values.

For this state-level data, we can use a GeoJSON file created with US Census Bureau polygons and ACS data that is available for download here.
- Subset only the 48 continental states to make the data easier to map.
- Reproject the states to a projection that better represents size using st_transform() (EPSG 6580 US/Texas-Centric Lambert Conformal).
- Load the polygon file into a simple features (sf) data frame using st_read().
- Join the data table and polygons using merge().
- Draw the map using plot().
- The colorRampPalette() function creates a palette using the specified colors that are evocative of the characteristics being mapped.
library(sf)
states = st_read("2015-2019-acs-states.geojson")
states = states[!(states$ST %in% c("AK", "HI")),]
states = st_transform(states, 6580)
states.weight= merge(states, weight.states[,c("ST","AVGWEIGHT")])
palette = colorRampPalette(c("navy", "gray", "red"))
plot(states.weight["AVGWEIGHT"], breaks="quantile", pal=palette)
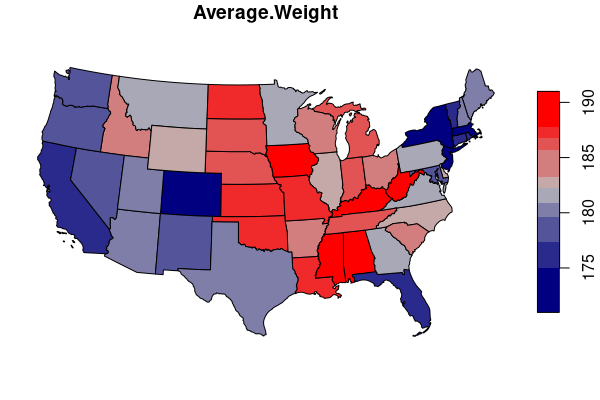
Mapping Proportions
For categorical variables, the most straighforward visualization will be a map of the percentage of one of the categories.
To calculate state-level proportions, first aggregate a table of sums of weights for the selected values, then divide by the sum of the weights to get a weighted proportion.
For this example, we get a weighted percent of respondents who have answered yes (1) to the question, "Ever told you had a heart attack, also called a myocardial infarction?
heart = responses[!is.na(responses$X_AAWEIGHT),]
heart$HEART.SUM = ifelse(heart$CVDINFR4 %in% 1, heart$X_AAWEIGHT, 0)
heart.states = aggregate(heart[,c("HEART.SUM", "X_AAWEIGHT")], by=list(ST = heart$ST), FUN=sum)
heart.states$HEART.ATTACK = 100 * heart.states$HEART.SUM / heart.states$X_AAWEIGHT
heart.states = heart.states[order(heart.states$HEART.ATTACK),]
head(heart.states)
ST HEART.SUM X_AAWEIGHT Heart.Attacks 8 DC 10365.90 768709.9 1.348480 31 NH 30773.18 1620881.6 1.898546 12 HI 25119.16 1202235.3 2.089371 7 CT 69825.33 3295819.6 2.118603 29 ND 17334.65 808834.5 2.143164 48 WA 154142.18 7009006.9 2.199201
library(sf)
states = st_read("2015-2019-acs-states.geojson")
states = states[!(states$ST %in% c("AK", "HI")),]
states = st_transform(states, 6580)
states.heart = merge(states, heart.states[,c("ST","HEART.ATTACK")])
palette = colorRampPalette(c("navy", "gray", "red"))
plot(states.heart["HEART.ATTACK"], breaks="quantile", pal=palette)
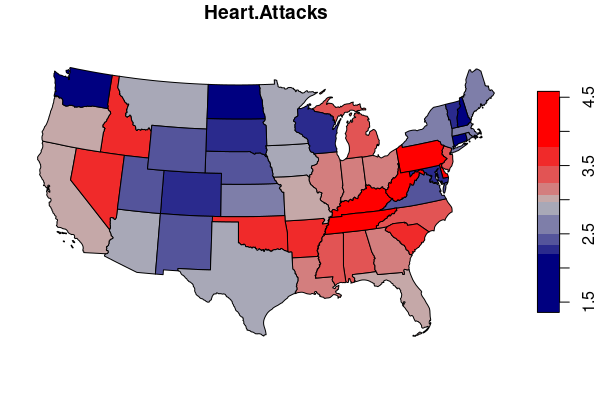
Classification
The choice of what colors to use for maps and what values to associate with those colors is both an aesthetic decision and a determinant of how readers will interpret a map.
With thematic choropleth maps like those used above, classification is the technique used to associate ranges of values with specific colors from the color palette used on the map. Classification choices are critical to the "story" a map will tell, and can be used to "lie" with maps (Monmonier 2018). Far from being objective representations of reality, maps incorporate numerous cartographic choices that reflect the subjectivities and goals of the cartographer.
With the plot() function from the sf library, the breaks parameter can be used to specify the algorithm for choosing the breaks between ranges of values associated with each color. The choice of algorithm will be influenced by the type of distribution. Some of the many options include:
- "equal" divides the range evenly. This creates maps where specific values for areas can be more-clearly interpreted, but can reduce contrast between areas when the breaks have no relationship to the clusters of values in the data. The "pretty" equal classification scheme is the default in the simple features plot() function.
- "jenks" is an algorithm that "seeks to reduce the variance within classes and maximize the variance between classes." It is a good general purpose algorithm commonly used by commercial GIS mapping software. The algorithm was originally proposed by cartographer George Jenks in 1967 (Wikipedia 2020).
- "quantile" creates quantile classes that have an equal number of values in each class. This will create a map that maximizes the variation in color. This may be useful with variables that have a uniform distribution, but can overemphasize small differences in values.
- "sd" will classify based on mean and standard deviation. This can be a good choice if you specifically want to emphasize deviation from the mean in unskewed, normally-distributed data.
par(mfrow=c(2,2))
palette = colorRampPalette(c("navy", "lightgray", "red"))
plot(states.heart["HEART.ATTACK"], pal=palette, breaks="fisher", main="Fisher-Jenks")
plot(states.heart["HEART.ATTACK"], pal=palette, breaks="quantile", main="Quantile")
plot(states.heart["HEART.ATTACK"], pal=palette, breaks="sd", main="Standard Deviation")
plot(states.heart["HEART.ATTACK"], pal=palette, breaks="equal", main="Equal Interval")
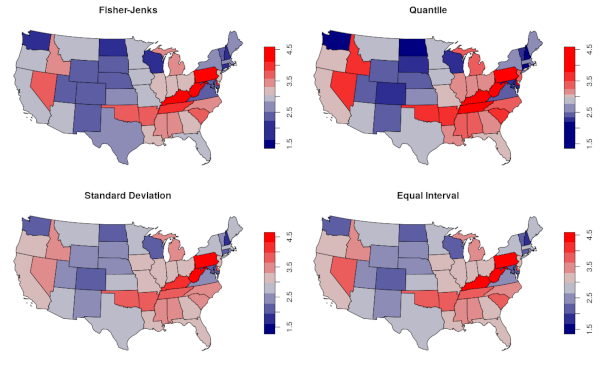
Modifiable Areal Unit Problem
Different choices of what types and boundaries of areas you used to aggregate spatial data can result in radically different results.
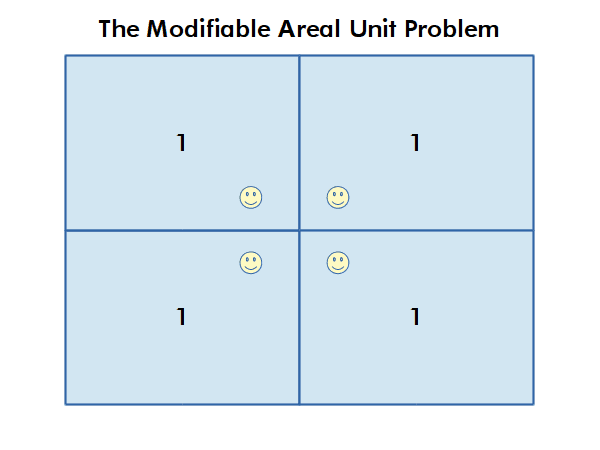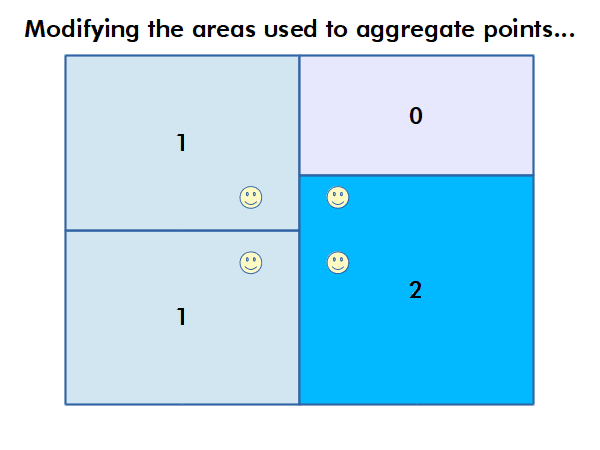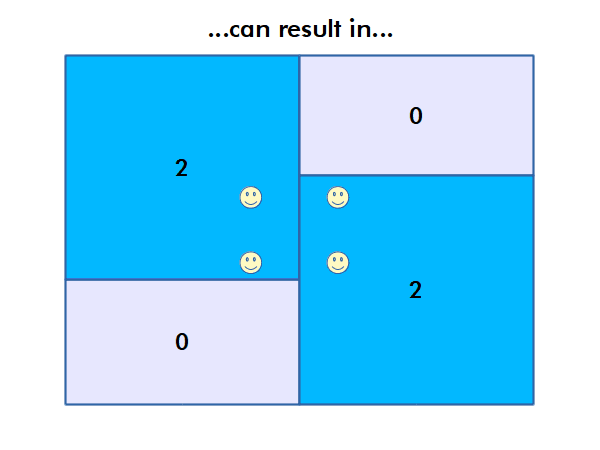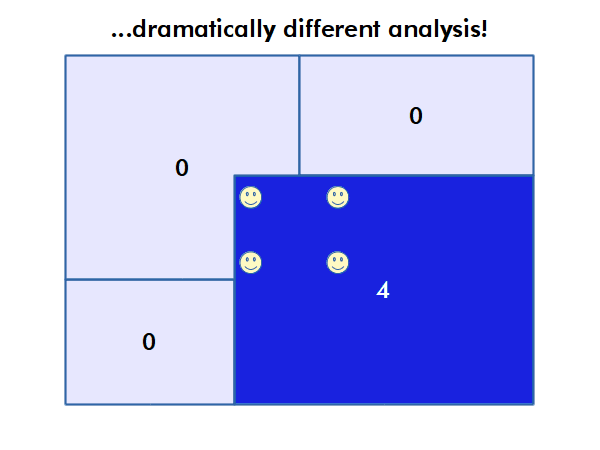
The modifiable areal unit problem (MAUP) is "a source of statistical bias that is common in spatially aggregated data, or data grouped into regions or districts where only summary statistics are produced within each district, especially where the districts chosen are not suitable for the data" (wiki.gis.com 2022).
For example, maps of electoral results give a very different impression depending on whether you map by state or by county.
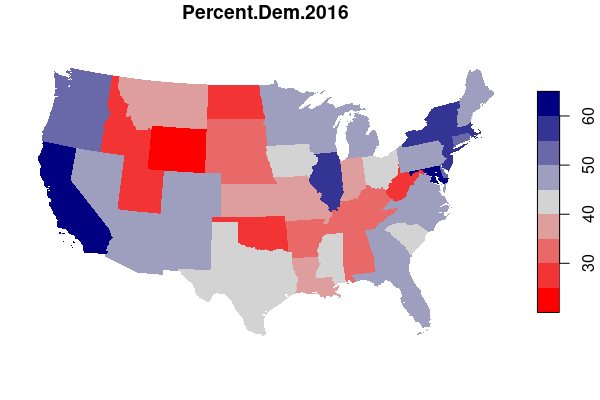
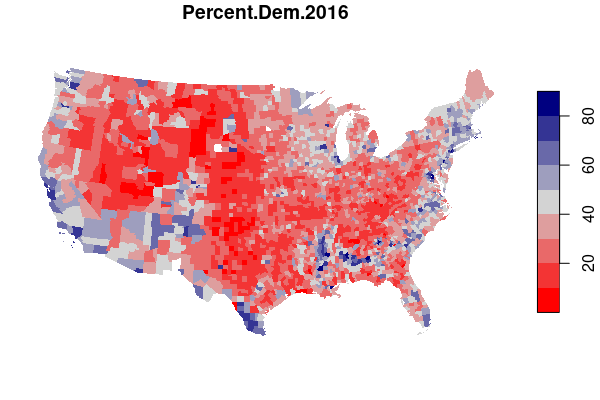
In drawing electoral boundaries, the MAUP is intentionally used in a process called gerrymandering, where specific groups of voters are either "packed" into a small number of homogeneous districts or "cracked" across multiple districts in order to reduce the power and representation available to those groups.
There is no universally applicable solution to the MAUP. In cases where accuracy is not essential (such as with data exploration) or possible (such as with data that has been aggregated for privacy reasons), the uncertainty presented by the MAUP may be acceptable as long as caveats are provided along with the analysis. In other cases, more-sophisticated analytical approaches may be more appropriate.
Selected Column Information
The following information on the variables selected above is from the BRFSS LLCP 2020 Codebook Report. If you use more-current data or other variables, you will want to consult the current codebook provided on the BRFSS website linked above.
- ST: Two-digit USPS state name abbreviation
- FIPS: Two-digit FIPS code
- X_URBSTAT:
- 1: urban county
- 2: rural county
- X_AGEG5YR: Age five year category
- 1: 18 - 24
- 2 - 13: Convert to start year with the formula 15 + (n * 5)
- 14: Don’t know/Refused/Missing
- DISPCODE: Final disposition
- 1100: Completed interview
- 1200: Partially completed interview
- NUMADULT: Number of adults in household
- SEXVAR: Sex of respondent
- 1: Male
- 2: Female
- GENHLTH: Would you say that in general your health is...
- 1 (excellent) to 5 (poor)
- 7: Don't know / not sure
- 9: Refused
- PHYSHLTH: Now thinking about your physical health, which includes physical
illness and injury, for how many days during the past 30 days was your physical
health not good? (number of days)
- 1 - 30: Number of days
- 88: None
- 77: Don’t know/Not sure
- 99: Refused
- NA: Not asked or Missing
- MENTHLTH:Now thinking about your mental health, which includes stress, depression, and problems with emotions, for how many days during the past 30 days was your mental health not good? (number of days)
- 1 - 30: Number of days
- 88: None
- 77: Don’t know/Not sure
- 99: Refused
- NA: Not asked or Missing
- HLTHPLN1: Do you have any kind of health care coverage, including health
insurance, prepaid plans such as HMOs, or government plans such as Medicare, or
Indian Health Service?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- EXERANY2: During the past month, other than your regular job, did you
participate in any physical activities or exercises such as running,
calisthenics, golf, gardening, or walking for exercise?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- SLEPTIM1: On average, how many hours of sleep do you get in a 24-hour period? (avg hours)
- 1 - 24: Number of hours
- 77: Don’t know/Not Sure
- 99: Refused
- NA: Missing
- CVDINFR4: Ever told you had a heart attack, also called a myocardial infarction?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- CVDCRHD4: Ever told you had angina or coronary heart disease?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- CVDSTRK3: (Ever told) (you had) a stroke?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- ASTHMA3: (Ever told) (you had) asthma?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- CHCSCNCR: (Ever told) (you had) skin cancer?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- CHCOCNCR: (Ever told) (you had) any other types of cancer?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- CHCCOPD2: (Ever told) (you had) chronic obstructive pulmonary disease, C.O.P.D., emphysema or chronic bronchitis?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- HAVARTH4: (Ever told) (you had) some form of arthritis, rheumatoid
arthritis, gout, lupus, or fibromyalgia? (Arthritis diagnoses include:
rheumatism, polymyalgia rheumatica; osteoarthritis (not osteporosis);
tendonitis, bursitis, bunion, tennis elbow; carpal tunnel syndrome, tarsal
tunnel syndrome; joint infection, etc.)
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- ADDEPEV3: (Ever told) (you had) a depressive disorder (including
depression, major depression, dysthymia, or minor depression)?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- CHCKDNY2: Not including kidney stones, bladder infection or incontinence,
were you ever told you had kidney disease?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- DIABETE4: (Ever told) (you had) diabetes?
- 1: Yes
- 2: Yes when pregnant
- 3: No
- 4: No but pre-diabetic or borderline
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- DIABAGE3: How old were you when you were told you had diabetes?
- 1 - 96: Age in years
- 97: 97 years of age or older
- 98: Don't know / Not sure
- 99: Refused
- NA: Not asked or missing
- LASTDEN4: Including all types of dentists, such as orthodontists, oral
surgeons, and all other dental specialists, as well as dental hygienists, how
long has it been since you last visited a dentist or a dental clinic for any
reason?
- 1: Within the past year (anytime less than 12 months ago)
- 2: Within the past 2 years (1 year but less than 2 years ago)
- 3: Within the past 5 years (2 years but less than 5 years ago)
- 4: 5 or more years ago
- 7: Don’t know/Not sure
- 8: Never
- 9: Refused
- NA: Missing
- RMVTETH4: Not including teeth lost for injury or orthodontics, how many of your permanent teeth have been removed because of tooth decay or gum disease? (1 one to five, 2 six or more, 3 all, 8 none)
- 1: 1 to 5
- 2: 6 or more, but not all
- 3: All
- 7: Don’t know/Not sure
- 8: None
- 9: Refused
- NA: Not asked or Missing
- MARITAL: Are you (marital status)
- 1: Married
- 2: Divorced
- 3: Widowed
- 4: Separated
- 5: Never married
- 6: A member of an unmarried couple
- 9: Refused
- NA: Not asked or Missing
- EDUCA: What is the highest grade or year of school you completed?
- 1: Never attended school or only kindergarten
- 2: Grades 1 through 8 (Elementary)
- 3: Grades 9 through 11 (Some high school)
- 4: Grade 12 or GED (High school graduate)
- 5: College 1 year to 3 years (Some college or technical school)
- 6: College 4 years or more (College graduate)
- 9: Refused
- NA: Not asked or Missing
- RENTHOM1: Do you own or rent your home?
- 1: Own
- 2: Rent
- 3: Other arrangement
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- VETERAN3: Have you ever served on active duty in the United States Armed
Forces, either in the regular military or in a National Guard or military
reserve unit?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- EMPLOY1: Are you currently...?
- 1: Employed for wages
- 2: Self-employed
- 3: Out of work for 1 year or more
- 4: Out of work for less than 1 year
- 5: A homemaker
- 6: A student
- 7: Retired
- 8: Unable to work
- 9: Refused
- NA: Not asked or Missing
- CHILDREN: How many children less than 18 years of age live in your household?
- 1 - 87: Number of children
- 88: None
- 99: Refused
- NA: Not asked or Missing
- INCOME2: Is your annual household income from all sources?
- 1: Less than $10,000
- 2: $10,000 to less than $15,000
- 3: $15,000 to less than $20,000
- 4: $20,000 to less than $25,000
- 5: $25,000 to less than $35,000
- 6: $35,000 to less than $50,000
- 7: $50,000 to less than $75,000)
- 8: $75,000 or more
- 77: Don’t know/Not sure
- 99: Refused
- NA: Not asked or Missing
- PREGNANT: To your knowledge, are you now pregnant?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- WEIGHT2: About how much do you weigh without shoes?
- 50 - 776: Weight (pounds)
- 7777: Don’t know/Not sure
- 9023 - 9352 Weight (kilograms)
- 9999:Refused
- NA: Not asked or Missing
- HTIN4: About how tall are you without shoes?
- 36 - 95: Height in inches
- NA: Don’t know/Refused/Not asked or Missing
- DEAF: Are you deaf or do you have serious difficulty hearing?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- BLIND: Are you blind or do you have serious difficulty seeing, even when wearing glasses?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- SMOKDAY2: Do you now smoke cigarettes every day, some days, or not at all?
- 1: Current smoker - now smokes every day
- 2: Current smoker - now smokes some days
- 3: Not at all
- 7: Don't know
- 9: Refused
- NA: Not asked or missing
- STOPSMK2: During the past 12 months, have you stopped smoking for one day
or longer because you were trying to quit smoking?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- AVEDRNK3: One drink is equivalent to a 12-ounce beer, a 5-ounce glass of wine, or a drink with one shot of liquor. During the past 30 days, on the days when you drank, about how many drinks did you drink on the average? (A 40 ounce beer would count as 3 drinks, or a cocktail drink with 2 shots would count as 2 drinks.)
- 1 - 76: Number of drinks
- 88: None
- 77: Don’t know/Not sure
- 99: Refused
- NA: Not asked or Missing
- DRNK3GE5: Considering all types of alcoholic beverages, how many times
during the past 30 days did you have 5 or more drinks for men or 4 or more
drinks for women on an occasion? (binge drinking in past 30 days)
- 1 - 76: Number of times
- 88: None
- 77: Don’t know/Not Sure
- 99: Refused
- NA: Not asked or Missing
- SEATBELT: How often do you use seat belts when you drive or ride in a car?
Would you say—
- 1: Always
- 2: Nearly always
- 3: Sometimes
- 4: Seldom
- 5: Never
- 7: Don’t know/Not sure
- 8: Never drive or ride in a car
- 9: Refused
- NA: Not asked or Missing
- HADMAM: Have you ever had a mammogram?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- PSATEST1: Have you ever had a P.S.A. test?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- COLNSCPY: Have you ever had a colonoscopy?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- HIVTST7: Including fluid testing from your mouth, but not including tests
you may have had for blood donation, have you ever been tested for H.I.V?
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- HIVRISK5: I am going to read you a list. When I am done, please tell me if
any of the situations apply to you. You do not need to tell me which one. You
have injected any drug other than those prescribed for you in the past year.
You have been treated for a sexually transmitted disease or STD in the past
year. You have given or received money or drugs in exchange for sex in the past
year.
- 1: Yes
- 2: No
- 7: Don’t know/Not Sure
- 9: Refused
- NA: Not asked or Missing
- X_LLCPWT (landline, cellphone weighting): This is a weighting factor that can be assigned to each response to compensate for uneven geographic coverage of survey administration (noncoverage) and lack of responsiveness from some segments of the population (nonresponse). See Weighting the BRFSS Data (CDC 2020) for more details.